□ Unsupervised Discovery of Temporal Structure in Noisy Data with Dynamical Components Analysis
>> https://arxiv.org/pdf/1905.09944v1.pdf
DCA robustly extracts dynamical structure in noisy, high-dimensional time series data while retaining the computational efficiency and geometric interpretability of linear dimensionality reduction methods.
Dynamical Components Analysis (DCA), a linear dimensionality reduction method which discovers a subspace of high-dimensional time series data with maximal predictive information, defined as the mutual information between the past and future.
Both the time- and frequency-domain implementations of DCA may be made differentiable in the input data, opening the door to extensions of DCA that learn nonlinear transformations of the input data, including kernel-like dimensionality expansion, or that use a nonlinear mapping from the high- to low-dimensional space, including deep architectures.

□ On the local and boundary behavior of mappings on factor-spaces
>> https://arxiv.org/pdf/1905.06414v1.pdf
the Poincar ́e theorem on uniformization, according to which each Riemannian surface is conformally equivalent to a certain factor-space of a flat domain with respect to the group of linear fractional mappings.
to establish modular inequalities on orbit spaces, and with their help to study the local and boundary behavior of maps with branching of arbitrary dimension, which are defined only in a certain domain and can have an unbounded quasi- conformality coefficient.
The map acting between domains of two factor spaces by certain groups of Mo ̈bius automorphisms.


□ Complete deconvolution of cellular mixtures based on linearity of transcriptional signatures
>> https://www.nature.com/articles/s41467-019-09990-5
identify a previously unrecognized property of tissue-specific genes – their mutual linearity – and use it to reveal the structure of the topological space of mixed transcriptional profiles and provide a noise-robust approach to the complete deconvolution problem.
Mathematically, non-zero singular vectors beyond the number of cell types arise because SVD attempts to fit the non-linear variation with linear components which are not relevant for the complete deconvolution procedure.
understanding the linear structure of the space revealed a major underappreciated aspect of both partial and complete deconvolution approaches: individual cell types often have varying cell size which leads to a limitation in identifying cellular frequencies.

□ Gauge Equivariant Convolutional Networks and the Icosahedral Convolutional Ceural Networks
>> https://arxiv.org/pdf/1902.04615.pdf
Vector fields don’t need to have the same dimension as the tangent space. Instead, they can have their own vector space of arbitrary dimension at each point.
the search for a geometrically natural definition of “manifold convolution”, a key problem in geometric deep learning, leads inevitably to gauge equivariance.
implement gauge equivariant CNNs for signals defined on the surface of the icosahedron, which provides a reasonable approximation of the sphere.
the general theory of gauge equivariant convolutional networks on manifolds, and demonstrated their utility in a special case: learning with spherical signals using the icosahedral convolutional neural network.

□ Reconstructing wells from high density regions extracted from super-resolution single particle trajectories
>> https://www.biorxiv.org/content/biorxiv/early/2019/05/20/642744.full.pdf
The biophysical properties of these regions are characterized by a drift and their extension (a basin of attraction) that can be estimated from an ensemble of trajectories.
two statistical methods to recover the dynamics and local potential wells (field of force and boundary) using as a model a truncated Ornstein-Ulhenbeck process.

□ APEC: An accesson-based method for single-cell chromatin accessibility analysis
>> https://www.biorxiv.org/content/biorxiv/early/2019/05/23/646331.full.pdf
an accessibility pattern-based epigenomic clustering (APEC) method, which classifies each individual cell by groups of accessible regions with synergistic signal patterns termed “accessons”.
a fluorescent tagmentation- and FACS-sorting-based single-cell ATAC-seq technique named ftATAC-seq and investigated the per cell regulome dynamics.
APEC also identifies significant differentially accessible sites, predicts enriched motifs, and projects pseudotime trajectories.

□ SAME-clustering: Single-cell Aggregated Clustering via Mixture Model Ensemble
>> https://www.biorxiv.org/content/biorxiv/early/2019/05/24/645820.full.pdf
SAME-clustering, a mixture model-based approach that takes clustering solutions from multiple methods and selects a maximally diverse subset to produce an improved ensemble solution.
In the current implementation of SAME-clustering, we first input a gene expression matrix into five individual clustering methods, SC3, CIDR, Seurat, t-SNE + k-means, and SIMLR, to obtain five sets of clustering solutions.
SAME-clustering assumes that these labels are drawn from a mixture of multivariate multinomial distributions to build an ensemble solution by solving a maximum likelihood problem using the expectation-maximization (EM) algorithm.

□ Benchmarking algorithms for gene regulatory network inference from single-cell transcriptomic data
>> https://www.biorxiv.org/content/biorxiv/early/2019/05/20/642926.full.pdf
the accuracy of the algorithms measured in terms of AUROC and AUPRC was moderate, by and large, although the methods were better in recovering interactions in the artificial networks than the Boolean models.
Techniques that did not require pseudotime-ordered cells were more accurate, in general. There were an excess of feed-forward loops in predicted networks than in the Boolean models.

□ Knowledge-guided analysis of 'omics' data using the KnowEnG cloud platform
>> https://www.biorxiv.org/content/biorxiv/early/2019/05/19/642124.full.pdf
The system offers ‘knowledge-guided’ data-mining and machine learning algorithms, where user-provided data are analyzed in light of prior information about genes, aggregated from numerous knowledge-bases and encoded in a massive ‘Knowledge Network’.
KnowEnG adheres to ‘FAIR’ principles: its tools are easily portable to diverse computing environments, run on the cloud for scalable and cost-effective execution of compute-intensive and data-intensive algorithms, and are interoperable with other computing platforms.
□ Illumination depth
>> https://arxiv.org/pdf/1905.04119v1.pdf
The concept of illumination bodies studied in convex geometry is used to amend the halfspace depth for multivariate data.
The illumination is, in a certain sense, dual to the halfspace depth mapping, and shares the majority of its beneficial properties. It is affine invariant, robust, uniformly consistent, and aligns well with common probability distributions.
The proposed notion of illumination enables finer resolution of the sample points, naturally breaks ties in the associated depth-based ordering, and introduces a depth-like function for points outside the convex hull of the support of the probability measure.
□ MetaQUBIC: a computational pipeline for gene-level functional profiling of metagenome and metatranscriptome
>> https://academic.oup.com/bioinformatics/advance-article-abstract/doi/10.1093/bioinformatics/btz414/5497255
MetaQUBIC, an integrated biclustering-based computational pipeline for gene module detection that integrates both metagenomic and metatranscriptomic data.
MetaQUBIC investigates 735 paired DNA and RNA samples, resulting in a comprehensive hybrid gene expression matrix of 2.3 million cross-species genes, and mapping datasets to the IGC reference database were proceeded on the XSEDE PSC cluster.
□ A hypergraph-based method for large-scale dynamic correlation study at the transcriptomic scale
>> https://bmcgenomics.biomedcentral.com/articles/10.1186/s12864-019-5787-x
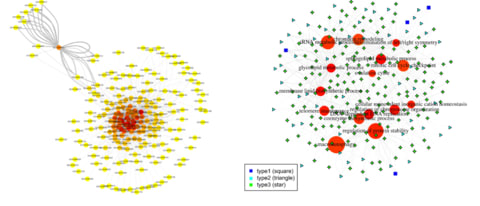
Hypergraph for Dynamic Correlation (HDC), to construct module-level three-way interaction networks.
The method is able to present integrative uniform hypergraphs to reflect the global dynamic correlation pattern in the biological system, providing guidance to down-stream gene triplet-level analyses.
□ SeRenDIP: SEquential REmasteriNg to DerIve Profiles for fast and accurate predictions of PPI interface positions
>> https://academic.oup.com/bioinformatics/advance-article-abstract/doi/10.1093/bioinformatics/btz428/5497259
With the aim of accelerating previous approach, they obtained sequence conservation profiles by re-mastering the alignment of homologous sequences found by PSI-BLAST.
SeRenDIP, SEquence-based Random forest predictor with lENgth and Dynamics for Interacting Proteins server offers a simple interface to our random-forest based method for predicting protein-protein interface positions from a single input sequence.

□ SciBet: An ultra-fast classifier for cell type identification using single cell RNA sequencing data
>> https://www.biorxiv.org/content/biorxiv/early/2019/05/23/645358.full.pdf
SciBet (Single Cell Identifier Based on Entropy Test), a Bayesian classifier that accurately predicts cell identity for any randomly sequenced cell.
SciBet addresses an important need in the rapidly evolving field of single- cell transcriptomics, i.e., to accurately and rapidly capture main features of diverse datasets regardless of technical factors or batch effect.
□ GeneEE: A universal method for gene expression engineering https://www.biorxiv.org/content/biorxiv/early/2019/05/23/644989.full.pdf
GeneEE, a straightforward method for generating artificial gene expression systems. GeneEE segments, contains a 200 nucleotide DNA with random nucleotide composition, can facilitate constitutive and inducible gene expression.
a DNA segment with random nucleotide composition can be used to generate artificial gene expression systems in seven different microorganisms.

□ PLASMA: Allele-Specific QTL Fine-Mapping
>> https://www.biorxiv.org/content/biorxiv/early/2019/05/25/650242.full.pdf
PLASMA is a novel, LD-aware method that integrates QTL and asQTL information to fine-map causal regulatory variants while drawing power from both the number of individuals and the number of allelic reads per individual.
PLASMA approach in combining QTL and AS signals opens up possible future work in two distinct directions. Generalizing from two to multiple phenotypes would be straightforward, and could utilize the colocalization algorithm first introduced in eCAVIAR.
□ Dynamic mode decomposition for analytic maps
>> https://arxiv.org/pdf/1905.09266v1.pdf
In the classical situations the reduction of the description to effective degrees of freedom has resulted in the derivation of transport equations for systems far from equilibrium using projection operator techniques,
an understanding of how dissipation emerges in many particle Hamiltonian systems, or the Bandtlow-Coveney equation for transport properties in discrete-time dynamical systems.
the modes identified by Extended dynamic mode decomposition correspond to those of compact Perron-Frobenius and Koopman operators defined on suitable Hardy-Hilbert spaces when the method is applied to classes of analytic maps.
□ Asymptotic behavior of the nonlinear Schrödinger equation on complete Riemannian manifold (R^n, g)
>> https://arxiv.org/pdf/1905.09540v1.pdf
Morawetz estimates for the system are directly derived from the metric g and are independent on the assumption of an Euclidean metric at infinity and the non-trapping assumption.
not only prove exponential stabilization of the system with a dissipation effective on a neighborhood of the infinity, but also prove exponential stabilization of the system with a dissipation effective outside of an unbounded domain.

□ Visualising quantum effective action calculations in zero dimensions
>> https://arxiv.org/pdf/1905.09674.pdf
an explicit treatment of the two-particle-irreducible (2PI) effective action for a zero-dimensional field theory.
the convexity of the 2PI effective action provides a comprehensive explanation of how the Maxwell construction arises in the case of multiple,f inding results that are consistent with previous studies of the one-particle-irreducible (1PI) effective action.
□ Anomalies in the Space of Coupling Constants and Their Dynamical Applications I
>> https://arxiv.org/abs/1905.09315
Failure of gauge invariance of the partition function under gauge transformations of these fields reflects ’t Hooft anomalies, the ordinary (scalar) coupling constants as background fields, i.e. to study the theory when they are spacetime dependent.
these anomalies and their applications in simple pedagogical examples in one dimension (quantum mechanics) and in some two, three, and four-dimensional quantum field theories.
An anomaly is an example of an invertible field theory, which can be described as an object in differential cohomology. an introduction to this perspective, and use Quillen’s superconnections to derive the anomaly for a free spinor field with variable mass.

□ Accelerating Sequence Alignment to Graphs
>> https://www.biorxiv.org/content/biorxiv/early/2019/05/27/651638.full.pdf
Given a variation graph in the form of a directed acyclic string graph, the sequence to graph alignment problem seeks to find the best matching path in the graph for an input query sequence.
Solving this problem exactly using a sequential dynamic programming algorithm takes quadratic time in terms of the graph size and query length, making it difficult to scale to high throughput DNA sequencing data.
the first parallel algorithm for computing sequence to graph alignments that leverages multiple cores and single-instruction multiple-data (SIMD) operations.
take advantage of the available inter-task parallelism, and provide a novel blocked approach to compute the score matrix while ensuring high memory locality.
Using a 48-core Intel Xeon Skylake processor, the proposed algorithm achieves peak performance of 317 billion cell updates per second (GCUPS), and demonstrates near linear weak and strong scaling on up to 48 cores.
It delivers significant performance gains compared to existing algorithms, and results in run-time reduction from multiple days to 3 hours for the problem of optimally aligning high coverage long or short DNA reads to an MHC human variation graph containing 10 million vertices.

□ Reconstruction of networks with direct and indirect genetic effects
>> https://www.biorxiv.org/content/biorxiv/early/2019/05/27/646208.full.pdf
an alternative strategy, where genetic effects are formally included in the graph. Using simulations, real data and statistical results show that this has important advantages:
genetic effects can be directly incorporated in causal inference, leading to the PCgen algorithm, which can handle many more traits than current approaches; and can test the existence of direct genetic effects, and also improve the orientation of edges between traits.

□ PhenoGeneRanker: A Tool for Gene Prioritization Using Complete Multiplex Heterogeneous Networks
>> https://www.biorxiv.org/content/biorxiv/early/2019/05/27/651000.full.pdf
PhenoGeneRanker, an improved version of a recently developed network propagation method called Random Walk with Restart on Multiplex Heterogeneous Networks (RWR-MH).
PhenoGeneRanker allows multi-layer gene and disease networks, and using using multi-omics datasets of rice to effectively prioritize the cold tolerance-related genes.
□ PSI-Sigma: a comprehensive splicing-detection method for short-read and long-read RNA-seq analysis
>> https://academic.oup.com/bioinformatics/advance-article-abstract/doi/10.1093/bioinformatics/btz438/5499131
PSI-Sigma, which uses a new PSI index (PSIΣ), and employed actual (non-simulated) RNA-seq data from spliced synthetic genes (RNA Sequins) to benchmark its performance (precision, recall, false positive rate, and correlation) in comparison with three leading tools.
PSI-Sigma outperformed these tools, especially in the case of AS events with multiple alternative exons and intron-retention events, and also briefly evaluated its performance in long-read RNA-seq analysis, by sequencing a mixture of human RNAs and RNA Sequins with nanopore long-read sequencers.

□ Linear time minimum segmentation enables scalable founder reconstruction
>> https://almob.biomedcentral.com/articles/10.1186/s13015-019-0147-6
a preprocessing routine relevant in pan-genomic analyses: consider a set of aligned haplotype sequences of complete human chromosomes.
Due to the enormous size of such data, one would like to represent this input set with a few founder sequences that retain as well as possible the contiguities of the original sequences.
an O(mn) time (i.e. linear time in the input size) algorithm to solve the minimum segmentation problem for founder reconstruction, improving over an earlier O(mn^2).
Given a segmentation S of R such that each segment induces exactly K distinct substrings, then construct a greedy parse P of R (and hence the corresponding set of founders) that has at most twice as many crossovers than the optimal parse in O (|S|×m) time and 𝑂(|S|×𝑚) space.

□ miRsyn: Identifying miRNA synergism using multiple-intervention causal inference
>> https://www.biorxiv.org/content/biorxiv/early/2019/05/28/652180.full.pdf
miRsyn is a novel framework called miRsyn for inferring miRNA synergism by using a causal inference method that mimics the effects in the multiple- intervention experiments, e.g. knock-down multiple miRNAs.
the identified miRNA synergistic network is small-world and biologically meaningful, and a number of miRNA synergistic modules are significantly enriched.

□ The Kipoi repository accelerates community exchange and reuse of predictive models for genomics
>> https://www.nature.com/articles/s41587-019-0140-0
Kipoi (Greek for ‘gardens’, pronounced ‘kípi’), an open science initiative to foster sharing and reuse of trained models in genomics.
Prominent examples include calling variants from whole-genome sequencing data, estimating CRISPR guide activity and predicting molecular phenotypes, including transcription factor binding, chromatin accessibility and splicing efficiency, from DNA sequence.
the Kipoi repository offers more than 2,000 individual trained models from 22 distinct studies that cover key predictive tasks in genomics, including the prediction of chromatin accessibility, transcription factor binding, and alternative splicing from DNA sequence.

□ KnockoffZoom: Multi-resolution localization of causal variants across the genome
>> https://www.biorxiv.org/content/biorxiv/early/2019/05/24/631390.full.pdf
KnockoffZoom, a flexible method for the genetic mapping of complex traits at multiple resolutions. KnockoffZoom localizes causal variants precisely and provably controls the false discovery rate using artificial genotypes as negative controls.
KnockoffZoom is equally valid for quantitative and binary phenotypes, making no assumptions about their genetic architectures. Instead, rely on well-established genetic models of linkage disequilibrium.
KnockoffZoom simultaneously addresses the current difficulties in locus discovery and fine-mapping by searching for causal variants over the entire genome and reporting the SNPs that appear to have a distinct influence on the trait while accounting for the effects of all others.
This work is facilitated by recent advances in statistics, notably knockoffs,23 whose general validity for GWAS has been explored and discussed before.
□ jackalope: a swift, versatile phylogenomic and high-throughput sequencing simulator
>> https://www.biorxiv.org/content/biorxiv/early/2019/05/27/650747.full.pdf
jackalope efficiently simulates variants from reference genomes and reads from both Illumina and PacBio platforms. Genomic variants can be simulated using phylogenies, gene trees, coalescent-simulation output, population-genomic summary statistics, and Variant Call Format files.
jackalope can simulate single, paired-end, or mate-pair Illumina reads, as well as reads from Pacific Biosciences. These simulations include sequencing errors, mapping qualities, multiplexing, and optical/PCR duplicates.
□ nf-core: Community curated bioinformatics pipelines
>> https://www.biorxiv.org/content/biorxiv/early/2019/05/26/610741.full.pdf
nf-core: a framework that provides a community-driven, peer- reviewed platform for the development of best practice analysis pipelines written in Nextflow.
Key obstacles in pipeline development such as portability, reproducibility, scalability and unified parallelism are inherently addressed by all nf-core pipelines.
□ trackViewer: a Bioconductor package for interactive and integrative visualization of multi-omics data
>> https://www.nature.com/articles/s41592-019-0430-y

□ CoCo: RNA-seq Read Assignment Correction for Nested Genes and Multimapped Reads
>> https://academic.oup.com/bioinformatics/advance-article/doi/10.1093/bioinformatics/btz433/5505419
CoCo uses a modified annotation file that highlights nested genes and proportionally distributes multimapped reads between repeated sequences.
as sequencing depth increases and the capacity to simultaneously detect both coding and non-coding RNA improves, read assignment tools like CoCo will become essential for any sequencing analysis pipeline.
□ AIVAR: Assessing concordance among human, in silico predictions and functional assays on genetic variant classification
>> https://academic.oup.com/bioinformatics/advance-article-abstract/doi/10.1093/bioinformatics/btz442/5505418
the results indicated that neural network model trained from functional assay data may not produce accurate prediction on known variants.
AIVAR (Artificial Intelligent VARiant classifier) was highly comparable to human experts on multiple verified data sets. Although highly accurate on known variants, AIVAR together with CADD and PhyloP showed non-significant concordance with SGE function scores.
□ CoMM-S2: a collaborative mixed model using summary statistics in transcriptome-wide association studies
>> https://www.biorxiv.org/content/biorxiv/early/2019/05/29/652263.full.pdf
a novel probabilistic model, CoMM-S2, to examine the mechanistic role that genetic variants play, by using only GWAS summary statistics instead of individual-level GWAS data.
an efficient variational Bayesian expectation-maximization accelerated using parameter expan- sion (PX-VBEM), where the calibrated evidence lower bound is used to conduct likelihood ratio tests for genome-wide gene associations with complex traits/diseases.
□ Smart computational exploration of stochastic gene regulatory network models using human-in-the-loop semi-supervised learning
>> https://academic.oup.com/bioinformatics/advance-article-abstract/doi/10.1093/bioinformatics/btz420/5505421
Discrete stochastic models of gene regulatory network models are indispensable tools for biological inquiry since they allow the modeler to predict how molecular interactions give rise to nonlinear system output.
Utilizing that similar simulation output is in proximity of each other in a feature space, the modeler can focus on informing the system about what behaviors are more interesting than others by labeling, rather than analyzing simulation results with custom scripts and workflows.
□ High-Dimensional Functional Factor Models
>> https://arxiv.org/pdf/1905.10325v1.pdf
This model and theory are developed in a general Hilbert space setting that allows panels mixing functional and scalar time series.
derive consistency results in the asymptotic regime where the number of series and the number of time observations diverge, thus exemplifying the "blessing of dimensionality" that explains the success of factor models in the context of high-dimensional scalar time series.